Our Focus
Ongoing research across a wide spectrum of biomedicine and the life sciences; including biochemistry, stem cell biology, immunology, molecular genetics, molecular medicine and gene therapy.
The Terry Fox Laboratory (TFL) has expertise in diverse research areas, including mammalian systems, normal and malignant mammary tissues, lineage specification in the gut, and yeast genetics. Experimental approaches employed at TFL include genetically engineered models, patient xenograft models, in vitro cell based assays, highly dimension single cell phenotyping, molecular biology and biochemistry, DNA regulation, transcriptomics, epigenetics, and DNA repair. In recent years, TFL has placed emphasis on the development of new technologies and their use to address fundamental questions in the control of cell growth and differentiation, aging, and gene regulation.
Blood Cancer
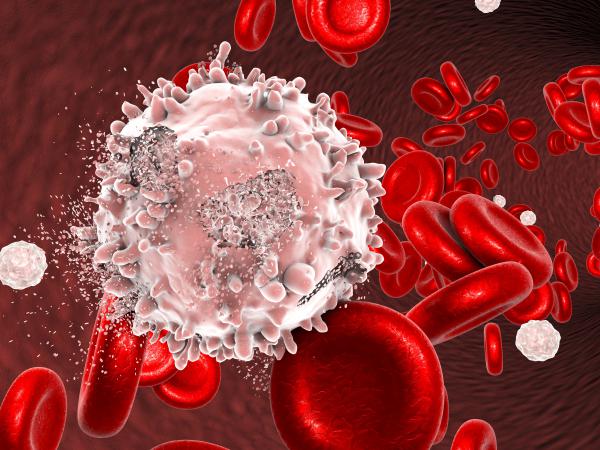
Working with innovative model systems and patients our research addresses the root causes of leukemias and myeloma and is developing new tools for their diagnosis, characterization and treatment.
Genome Dynamics

Genes influence every aspect of human biology and disease. We are studying how our genes and genomes are faithfully maintained as we age and the complex ways in which they are regulated to control development or are dysregulated during cancer formation.
Stem Cells
Stem cells are critical components in healthy aging and tumour suppression since they enable the development and maintenance of adult tissues. We are working to understand how stem cell properties enable blood, breast and liver development and how these properties are co-opted in cancers.
Immunology
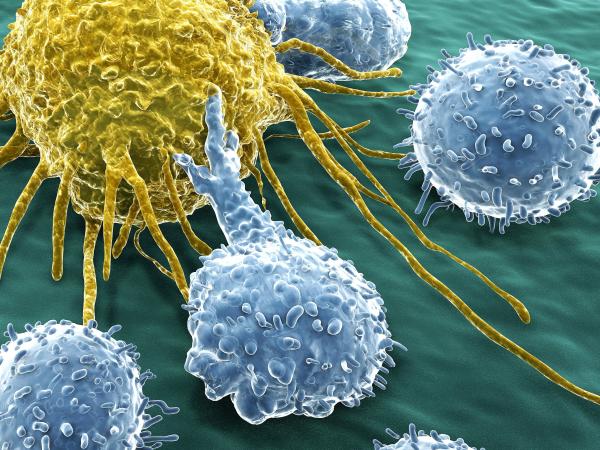
Immunology is the study of the immune system, including its functions, responses to pathogens and foreign substances, and how it protects against disease. We are investigating the interaction of the immune system with normal and cancer cells to develop novel immunotherapies to use the body’s own immune system to fight cancer.
Research Labs & Principal Investigators
Emeritus Scientists

Allen Eaves, PhD, FRCPC, FACP

R. Keith Humphries, MD, PhD, FRS(C)

Donna Hogge, MD, PhD

Dixie Mager, PhD

Ryan Brinkman, PhD

Gerald (Gerry) Krystal, PhD
Scientists In Memorial

Connie Eaves, PhD (1944-2024)
News & Events
Recent Publications
The ups and downs of maturing zonated hepatoctyes.
BC Cancer Foundation is the fundraising partner of BC Cancer, which includes BC Cancer Research. Together with our donors, we are changing cancer outcomes for British Columbians by funding innovative research and personalized treatment and care.














