Scientific Highlights
1. Developed B9, an IL-6-dependent cell line, instrumental for IL-6 research.
Following my training as MD, I worked for 2 years on a tumor immunology at the Erasmus University in Rotterdam, the Netherlands During this time I reproduced the hybridoma work described by Kohler and Milstein (1) for which these investigators got the Nobel Prize in 1984. Based on my hybridoma expertise, I was hired in 1979 as a research fellow to set up monoclonal antibody technology in the Central Laboratory of the Blood Transfusion Service in Amsterdam (now called Sanquin), one of the premier immunology laboratory in the Netherlands. There I developed assays to detect cell monoclonal antibodies to surface antigens (2,3) and made monoclonal antibodies, some of which were used in the International Workshops to define Clusters of Differentiation (CD) antigens. For this workshop we had to submit large amounts of antibodies, which was not a problem except for one hybridoma cell line that could only be expanded in the presence of tissue culture medium conditioned by endothelial cells (4). We later found that this hybridoma (B9) maintained a strict requirement for IL-6 for growth (5-7). This line has been instrumental in the IL-6 field ( >1200 citations). A Canadian twist to this story is that I could convince Jack Gauldie from McMaster University that the hepatocyte stimulating factor he was working on was most likely was IL-6, a prediction which turned out to be correct, resulting in a widely (>2000) cited paper (8) and US patent No. 4,973,478.
References:
- Kohler, G. and Milstein, C. (1975) Continuous cultures of fused cells secreting antibody of predefined specificity. Nature, 256, 495-497.
- Lansdorp, P.M., Astaldi, G.C., Oosterhof, F., Janssen, M.C. and Zeijlemaker, W.P. (1980) Immunoperoxidase procedures to detect monoclonal antibodies against cell surface antigens. Quantitation of binding and staining of individual cells. J Immunol Methods, 39, 393-405.
- Lansdorp, P.M., van der Kwast, T.H., de Boer, M. and Zeijlemaker, W.P. (1984) Stepwise amplified immunoperoxidase (PAP) staining. I. Cellular morphology in relation to membrane markers. J Histochem Cytochem, 32, 172-178.
- Astaldi, G.C., Janssen, M.C., Lansdorp, P., Willems, C., Zeijlemaker, W.P. and Oosterhof, F. (1980) Human endothelial culture supernatant (HECS): a growth factor for hybridomas. J Immunol, 125, 1411-1414.
- Aarden, L.A., De Groot, E.R., Schaap, O.L. and Lansdorp, P.M. (1987) Production of hybridoma growth factor by human monocytes. Eur J Immunol, 17, 1411-1416.
- Brakenhoff, J.P., de Groot, E.R., Evers, R.F., Pannekoek, H. and Aarden, L.A. (1987) Molecular cloning and expression of hybridoma growth factor in Escherichia coli. J Immunol, 139, 4116-4121.
- Lansdorp, P.M., Aarden, L.A., Calafat, J. and Zeiljemaker, W.P. (1986) A growth-factor dependent B-cell hybridoma. Curr Top Microbiol Immunol, 132, 105-113.
- Gauldie, J., Richards, C., Harnish, D., Lansdorp, P. and Baumann, H. (1987) Interferon beta 2/B-cell stimulatory factor type 2 shares identity with monocyte-derived hepatocyte-stimulating factor and regulates the major acute phase protein response in liver cells. Proc Natl Acad Sci U S A, 84, 7251-7255.
2. Discovered tetrameric antibody complexes, reagents that are key to the success of Vancouver's STEMCell Technologies.
In Amsterdam, I made mouse IgG1 monoclonal antibodies to peroxidase to link peroxidase to cell surface antigens via mouse IgG1 monoclonal antibodies against cell surface antigens using polyclonal anti-mouse IgG1 (1,2). Based on the bivalency of IgG antibodies and the presence of two identical heavy chains in each of such antibody molecules, I speculated that monoclonal antibodies specific for selected epitopes present on the heavy chain of mouse IgG1 molecules might be able to cross-link two mouse IgG1 monoclonal antibodies into a stable tetrameric antibody complex. To test this hypothesis, I made (and found!) suitable (rat) monoclonal antibodies specific for mouse IgG1 (3). See Figure 1.
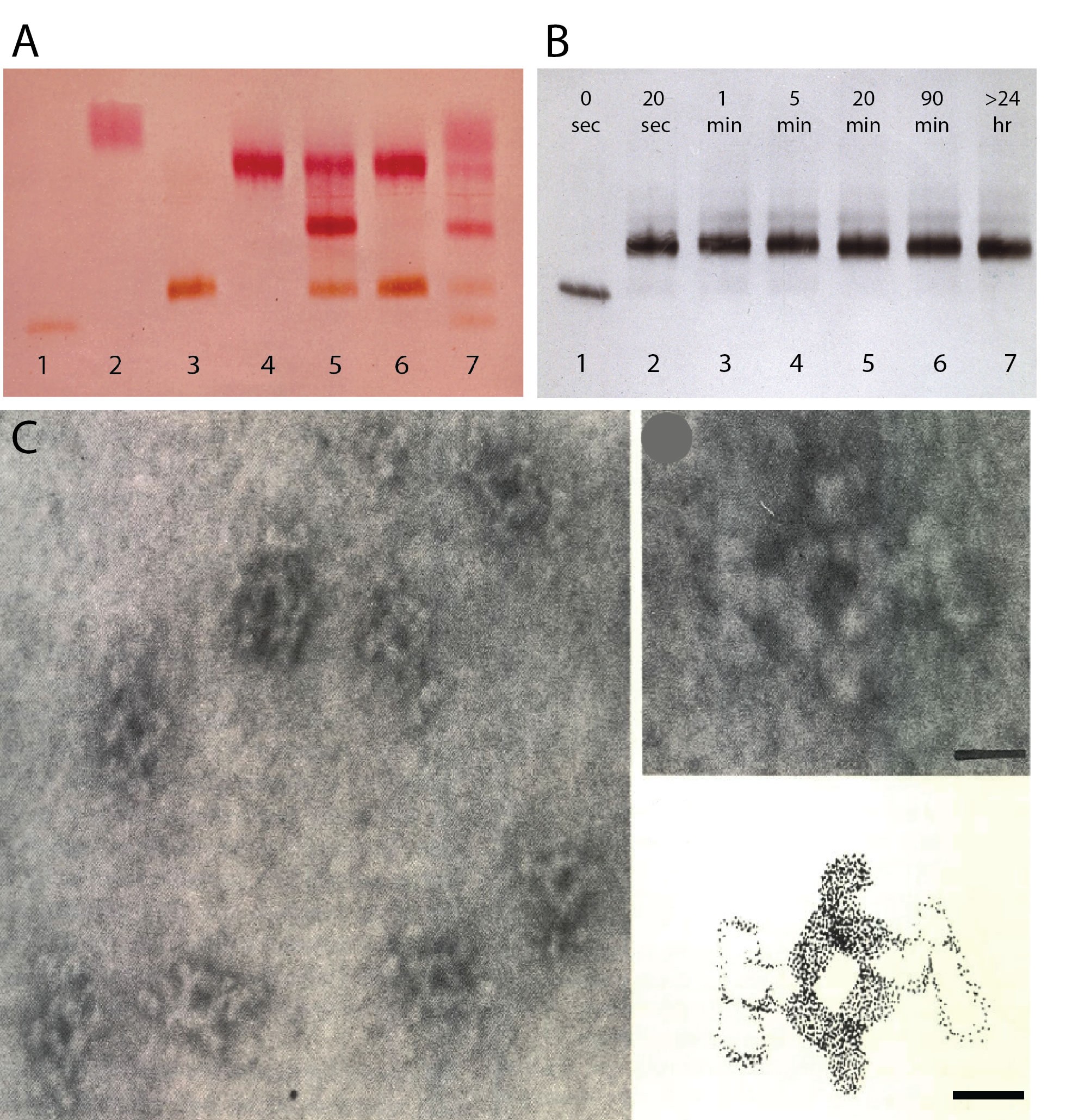
Figure 1. Tetrameric complexes of monoclonal rat and mouse antibodies (from (3). A. Staining for peroxidase and alkaline phosphatase following agar gel electrophoresis of mouse IgG1 anti-enzyme antibodies before and after addition of selected monoclonal rat anti-mouse IgG1. Lane 1: mouse IgG1 anti-peroxidase (a). Lane 2: mouse IgG1 anti- alkaline phosphatase (b). Lane 3: mouse IgG1 anti peroxidase plus an equimolar amount of rat anti-mouse IgG1. Note: no free mouse antibody. Lane 4: mouse IgG1 anti- alkaline phosphatase plus rat anti-mouse IgG1. Lane 5: equimolar amounts of mouse IgG1 anti- peroxidase and mouse IgG1 anti- alkaline phosphatase mixed before addition of an equimolar amount of rat anti-mouse IgG1. Note middle band with bispecific monoclonal tetrameric antibody complexes! Lane 6: A mixture of the complexes shown in lane 3 and 4. Note the absence of bispecific complexes, indicating high stability of tetrameric antibody complexes. Lane 7. As lane 5, using a limiting amount of rat anti-mouse IgG1 which shows free mouse IgG1 antibodies. B. Time course of tetrameric antibody complex formation. Mouse IgG1 anti-peroxidase (lane 1) was mixed with an equimolar amount of monoclonal rat anti-mouse IgG1. At the indicated time points the mixture was subjected to agarose gel electrophoresis and anti-peroxidase was visualized after blotting as in A. Note that antibody complexes are formed within seconds and not affected by electrophoresis. C. Scanning electron microscopy of purified tetramolecular antibody complexes. Scale bar 10 nm.
The ability to easily crosslink two different monoclonal antibodies into stable tetrameric antibody complexes (US patent No. 4,868,109) has found numerous applications in biotechnology. For example, we described applications in cell separation (4,5) and fluorescent labeling of cell surface antigens (6). Tetrameric antibody complexes are key to cell separation and many other technologies commercialized by STEMCELL Technologies in Vancouver, the largest biotechnology company in Canada.
References:
- Lansdorp, P.M., Astaldi, G.C., Oosterhof, F., Janssen, M.C. and Zeijlemaker, W.P. (1980) Immunoperoxidase procedures to detect monoclonal antibodies against cell surface antigens. Quantitation of binding and staining of individual cells. J Immunol Methods, 39, 393-405.
- Lansdorp, P.M., van der Kwast, T.H., de Boer, M. and Zeijlemaker, W.P. (1984) Stepwise amplified immunoperoxidase (PAP) staining. I. Cellular morphology in relation to membrane markers. J Histochem Cytochem, 32, 172-178.
- Lansdorp, P.M., Aalberse, R.C., Bos, R., Schutter, W.G. and Van Bruggen, E.F. (1986) Cyclic tetramolecular complexes of monoclonal antibodies: a new type of cross-linking reagent. Eur J Immunol, 16, 679-683.
- Thomas, T.E., Abraham, S.J., Otter, A.J., Blackmore, E.W. and Lansdorp, P.M. (1992) High gradient magnetic separation of cells on the basis of expression levels of cell surface antigens. J Immunol Methods, 154, 245-252.
- Thomas, T.E., Sutherland, H.J. and Lansdorp, P.M. (1989) Specific binding and release of cells from beads using cleavable tetrameric antibody complexes. J Immunol Methods, 120, 221-231.
- Wognum, A.W., Thomas, T.E. and Lansdorp, P.M. (1987) Use of tetrameric antibody complexes to stain cells for flow cytometry. Cytometry, 8, 366-371.
3. Produced several widely used monoclonal antibodies for human stem cell research (e.g. CD34, CD90).
During my graduate studies in Amsterdam my research interest shifted from immunology to human stem cell biology and the purification of blood-forming stem cells using monoclonal antibodies became a research priority for me. At the time, it was believed that purified stem cells, together with defined culture conditions, would allow unlimited expansion (self-renewal) of stem cells for clinical purposes, such as gene therapy and transplantation. During a meeting in London I met with Allen Eaves, the director of the Terry Fox Laboratory at the time. Allen convinced me to visit Vancouver and to do postdoctoral work in the lab of his wife, Connie Eaves, enabled by a Terry Fox fellowship. We made several monoclonal antibodies with remaining relevance for stem cell research including antibodies to CD34 and CD90. Our antibodies proved to be superior reagents to detect e.g. CD34 cells in peripheral blood (1) and identify and isolate stem cells in combination with antibodies to CD38 (2).
References:
- Siena, S., Bregni, M., Brando, B., Belli, N., Ravagnani, F., Gandola, L., Stern, A.C., Lansdorp, P.M., Bonadonna, G. and Gianni, A.M. (1991) Flow cytometry for clinical estimation of circulating hematopoietic progenitors for autologous transplantation in cancer patients. Blood, 77, 400-409.
- Terstappen, L.W., Huang, S., Safford, M., Lansdorp, P.M. and Loken, M.R. (1991) Sequential generations of hematopoietic colonies derived from single nonlineage-committed CD34+CD38- progenitor cells. Blood, 77, 1218-1227.
4. Developed serum-free culture medium and novel assays for hematopoietic stem cells.
Before the use of xeno-transplants, the activity of “candidate” hematopoietic stem cells was primarily measured by the ability of cells to produce colony-forming cells in long-term “Dexter” cultures (1). Together with Connie Eaves and Heather Sutherland we established a long-term culture assay for human cells (2) using the limiting dilution principles I used for producing monoclonal antibodies. This assay was used to purify “candidate” stem cells and for development of a serum-free culture medium for human “candidate” stem cells (3). This serum-free culture medium is currently available as “StemSPAN” from STEMCell technologies.
References:
- Dexter, T.M., Allen, T.D., Lajtha, L.G., Schofield, R. and Lord, B.I. (1973) Stimulation of differentiation and proliferation of haemopoietic cells in vitro. J Cell Physiol, 82, 461-473.
- Sutherland, H.J., Eaves, C.J., Eaves, A.C., Dragowska, W. and Lansdorp, P.M. (1989) Characterization and partial purification of human marrow cells capable of initiating long-term hematopoiesis in vitro. Blood, 74, 1563-1570.
- Mayani, H., Dragowska, W. and Lansdorp, P.M. (1993) Characterization of functionally distinct subpopulations of CD34+ cord blood cells in serum-free long-term cultures supplemented with hematopoietic cytokines. Blood, 82, 2664-2672.
5. Discovered that self-renewal properties of stem cells are developmentally controlled.
Despite our novel and highly specific monoclonal antibodies, innovative cell purification techniques, improved cell culture media and assays for stem cells, our attempts to increase the number of adult blood-forming stem cells in tissue culture were a dismal failure. It was possible to maintain CD34+ cells in culture for many weeks. However, we could not markedly increase the number of CD34+ cells when cultures were initiated with purified “candidate” stem cells from adult bone marrow (1). A breakthrough came when we finally tested “candidate” stem cells from fetal liver and umbilical cord blood in our culture systems. The number of CD34+ cells increased several thousand-fold in cultures of fetal liver cells and several hundred-fold in cord blood cultures. We concluded that the self-renewal of stem cells is developmentally controlled (2). Similar findings were obtained with purified murine stem cell “candidates” (3). Developmental changes in stem cell function were confirmed many years later (4,5).
References:
- Mayani, H., Dragowska, W. and Lansdorp, P.M. (1993) Lineage commitment in human hemopoiesis involves asymmetric cell division of multipotent progenitors and does not appear to be influenced by cytokines. J Cell Physiol, 157, 579-586.
- Lansdorp, P.M., Dragowska, W. and Mayani, H. (1993) Ontogeny-related changes in proliferative potential of human hematopoietic cells. J Exp Med, 178, 787-791.
- Rebel, V.I., Miller, C.L., Thornbury, G.R., Dragowska, W.H., Eaves, C.J. and Lansdorp, P.M. (1996) A comparison of long-term repopulating hematopoietic stem cells in fetal liver and adult bone marrow from the mouse. Exp Hematol, 24, 638-648.
- Kim, I., Saunders, T.L. and Morrison, S.J. (2007) Sox17 dependence distinguishes the transcriptional regulation of fetal from adult hematopoietic stem cells. Cell, 130, 470-483.
- Bowie, M.B., Kent, D.G., Dykstra, B., McKnight, K.D., McCaffrey, L., Hoodless, P.A. and Eaves, C.J. (2007) Identification of a new intrinsically timed developmental checkpoint that reprograms key hematopoietic stem cell properties. Proc Natl Acad Sci U S A, 104, 5878-5882.
6. Reported loss of telomeric DNA in human blood - forming stem cells.
The failure to significantly expand “candidate” hematopoietic stem cells from adult bone marrow in tissue culture was a set-back for therapeutic strategies counting on significant production of stem cells in tissue culture. The situation did not improve when we found that hematopoietic stem cells lost telomeric DNA with each cell divisions (1), similar to human fibroblasts (2).
Reference:
- Vaziri, H., Dragowska, W., Allsopp, R.C., Thomas, T.E., Harley, C.B. and Lansdorp, P.M. (1994) Evidence for a mitotic clock in human hematopoietic stem cells: loss of telomeric DNA with age. Proc Natl Acad Sci U S A,91, 9857-9860.
- Allsopp, R.C., Vaziri, H., Patterson, C., Goldstein, S., Younglai, E.V., Futcher, A.B., Greider, C.W. and Harley, C.B. (1992) Telomere length predicts replicative capacity of human fibroblasts. Proc Natl Acad Sci U S A,89, 10114-10118.
7. Developed quantitative fluorescence in situ hybridization (Q-FISH) techniques.
Questions about the role of telomeres in the biology of blood-forming stem cells were complicated by the laborious Southern analysis technique used to measure the length of telomere repeats requiring DNA from millions of cells. During a sabbatical in the Sylvius laboratory in Leiden, I explored using fluorescence in situ hybridization (FISH) techniques to measure the length of telomere repeats in chromosome spreads on microscope slides. The breakthrough came when I could get my hands on novel, synthetic “peptide nucleic acid” (PNA) probes (1). Briefly, I found that FISH techniques could be made quantitative (Q-FISH) using PNA probes under stringent, low ionic strength conditions, that allow PNA, but not competing single stranded DNA, to anneal to the selected target sequences. We published our findings (2) and obtained a US Patent (No 6,514,693).
References:
- Egholm, M., Buchardt, O., Christensen, L., Behrens, C., Freier, S.M., Driver, D.A., Berg, R.H., Kim, S.K., Norden, B. and Nielsen, P.E. (1993) PNA hybridizes to complementary oligonucleotides obeying the Watson-Crick hydrogen-bonding rules. Nature, 365, 566-568.
- Lansdorp, P.M., Verwoerd, N.P., van de Rijke, F.M., Dragowska, V., Little, M.T., Dirks, R.W., Raap, A.K. and Tanke, H.J. (1996) Heterogeneity in telomere length of human chromosomes. Hum Mol Genet, 5, 685-691.
8. Reported that without telomerase mice lose around 5kb of telomeric DNA per generation.
With PNA and Q-FISH tools, we contributed to a number of important studies, the most cited one being our contribution to the study of the telomerase knock-out (KO) KO mouse (1) (> 2000 citations). This paper was already under review at Cell when we got involved. Our findings significantly changed the message, essentially from “telomerase is not as important as we believed: it can be knocked out without consequences” to “telomerase KO mice lose 5 kb of telomeric DNA with each generation and uncapped telomeres at late generations cause chromosome fusions and genome instability”. See Figure.
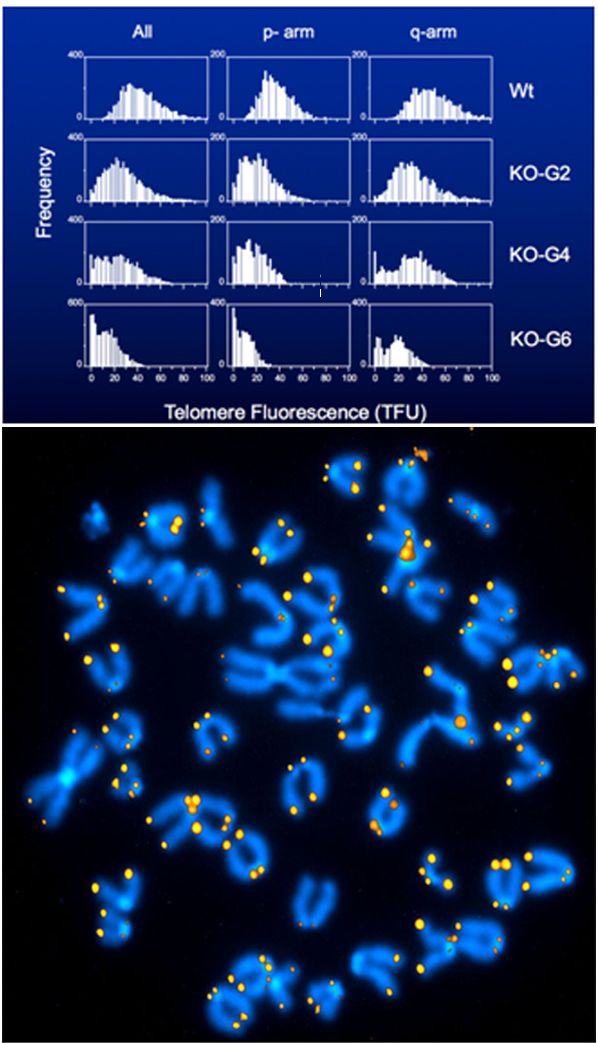
Figure. Telomerase KO mice lose about 5 kb of telomere repeats per generation. Q-FISH analysis of embryonic fibroblasts from the indicated subsequent generations of telomerase KO mice. 1 TFU corresponds to ~1kb of telomere repeats. Note that in wild-type (WT) animals all chromosomes are “capped” with readily detectable telomere repeats, whereas at generation 6 (bottom panel and chromosome spread) many chromosome ends have no telomere repeats. Note the dicentric chromosomes without telomere repeats at the point of fusion point (e.g. at 9 pm).
Reference:
- Blasco, M.A., Lee, H.W., Hande, M.P., Samper, E., Lansdorp, P.M., DePinho, R.A. and Greider, C.W. (1997) Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell, 91, 25-34.
9. Described variation in the length of telomere repeats at individual human chromosome ends.
We found that the average number of repeats of the telomere sequence (TTAGGG) on specific human chromosome arms is very similar in different tissues from the same donor but varies markedly between human donors (1). In all individuals studied, telomeres on chromosome 17p were shorter than the median telomere length, a finding that was confirmed by analysis of terminal restriction fragments from sorted chromosomes (2) and, many more years later, using a sequencing approach (3).
References:
- Martens, U.M., Zijlmans, J.M., Poon, S.S., Dragowska, W., Yui, J., Chavez, E.A., Ward, R.K. and Lansdorp, P.M. (1998) Short telomeres on human chromosome 17p. Nat Genet, 18, 76-80.
- Britt-Compton, B., Rowson, J., Locke, M., Mackenzie, I., Kipling, D. and Baird, D.M. (2006) Structural stability and chromosome-specific telomere length is governed by cis-acting determinants in humans. Hum Mol Genet, 15, 725-733.
- Karimian, K., Groot, A., Huso, V., Kahidi, R., Tan, K.T., Sholes, S., Keener, R., McDyer, J.F., Alder, J.K., Li, H. et al. (2024) Human telomere length is chromosome end-specific and conserved across individuals. Science, 384, 533-539.
10. Reported that a helicase-like gene is required to maintain guanine-rich DNA in C.elegans.
Intrigued by the large variation in the average length of telomere repeats at chromosome ends between cells of different individuals, I became intrigued by the genetic factors that regulate average telomere length. We initially focused on studies of mice in collaboration with Richard Hodes at NIH. In crosses between M.spretus (with short telomeres) and M.musculus (with long telomeres), a region on mouse chromosome 2 was identified that is required to elongate the short telomeres of M. spretus (1). However, at the time this region was too large (>10 Mb) to consider knocking out all the “candidate” genes in this region. Instead, we decided on further studies of a very similar gene in C.elegans, called “novel helicase-like” in collaboration with the lab of Ann Rose at UBC. We found that animals lacking the “novel helicase-like” gene displayed a striking genome instability phenotype: deletions throughout the genome that invariable started at the 3’ end of poly-guanine tracks longer than 18 nucleotides (2). We renamed the gene Dog 1 (for deletions of guanine rich-DNA) and we proposed that the DOG-1, the helicase encoded by the dog-1 gene could be required to resolve guanine quadruplex (G4) structures of G-rich DNA arising during replication. This pioneering work allowed us to deduce the function of a helicase protein from the signature phenotype observed in its absence. Later other groups showed that the homolog of dog-1 in humans (FANCJ) is indeed required to prevent instability at G-rich DNA (3,4).
References:
- Zhu, L., Hathcock, K.S., Hande, P., Lansdorp, P.M., Seldin, M.F. and Hodes, R.J. (1998) Telomere length regulation in mice is linked to a novel chromosome locus. Proc Natl Acad Sci U S A, 95, 8648-8653.
- Cheung, I., Schertzer, M., Rose, A. and Lansdorp, P.M. (2002) Disruption of dog-1 in Caenorhabditis elegans triggers deletions upstream of guanine-rich DNA. Nat Genet, 31, 405-409.
- Wu, Y., Shin-ya, K. and Brosh, R.M., Jr. (2008) FANCJ helicase defective in Fanconia anemia and breast cancer unwinds G-quadruplex DNA to defend genomic stability. Mol Cell Biol, 28, 4116-4128.
- London, T.B., Barber, L.J., Mosedale, G., Kelly, G.P., Balasubramanian, S., Hickson, I.D., Boulton, S.J. and Hiom, K. (2008) FANCJ is a structure-specific DNA helicase associated with the maintenance of genomic G/C tracts. J Biol Chem, 283, 36132-36139.
11. Cloned and described the function of RTEL1, a helicase required for maintaining telomeric DNA.
Encouraged by our findings in C.elegans we decided to knock-out “novel helicase-like” in the mouse in collaboration with Andras Nagy and Hao Ding in Toronto. We could demonstrate that this gene is indeed a major regulator of telomere length. We named the gene accordingly: Regulator of telomere length (Rtel) (1). This work has stood the test of time and patients with mutations in RTEL1 were described by several groups (2,3). Such patients have very short telomeres and often show bone marrow failure, similar to patients with mutations in telomerase genes such as TERT or TERC.
References:
- Ding, H., Schertzer, M., Wu, X., Gertsenstein, M., Selig, S., Kammori, M., Pourvali, R., Poon, S., Vulto, I., Chavez, E. et al. (2004) Regulation of murine telomere length by Rtel: an essential gene encoding a helicase-like protein. Cell, 117, 873-886.
- Walne, A.J., Vulliamy, T., Kirwan, M., Plagnol, V. and Dokal, I. (2013) Constitutional mutations in RTEL1 cause severe dyskeratosis congenita. Am J Hum Genet, 92, 448-453.
- Le Guen, T., Jullien, L., Touzot, F., Schertzer, M., Gaillard, L., Perderiset, M., Carpentier, W., Nitschke, P., Picard, C., Couillault, G. et al. (2013) Human RTEL1 deficiency causes Hoyeraal-Hreidarsson syndrome with short telomeres and genome instability. Hum Mol Genet, 22, 3239-3249.
12. Founded Repeat Diagnostics, a company that provides clinical measurements of the average length of telomere repeats at chromosome ends in nucleated blood cells.
We developed and perfected a method to measure the average length of telomeres in specific nucleated cell types in peripheral blood using PNA probes, fluorescence in situ hybridization (FISH) and flow cytometry, a technique we called flow FISH (1). With this method, we showed that there is a large variation in the average telomere length between individuals of the same age and that the most dramatic decline in telomere length occurs in the first few years of life (see Figure). Interestingly, at birth and throughout life the average telomere length was found to be longer in females than in males (2) and Figure, left panel). Based on reported sex differences in gene expression between cells from human embryos, I recently proposed that such telomere length differences could be established before embryo implantation (3). In collaboration with hematologists and geneticists at NIH, we established that the telomere length in patients with genetic defects in telomere pathway genes, such as TERT, TERC, RTEL1 and others could readily be distinguished from patients with other causes for their bone marrow failure (reviewed in (4). Because our flow FISH measurements could also distinguish carriers from telomerase mutations from siblings without such genetic defect (Figure, right panel), we obtained many requests from clinicians to perform diagnostic flow FISH experiments for their patients in our research lab. This clearly was not ideal, and I decided to set up a company, Repeat Diagnostics Inc., to provide clinical telomere length measurements. This company employs 8 people and is processing 10-20 samples per day for clinicians around the world looking after patients suspected of inherited or acquired “telomere biology disorders”. My take on the role of telomeres and telomerase in aging and cancer is summarized in two review papers (5,6).
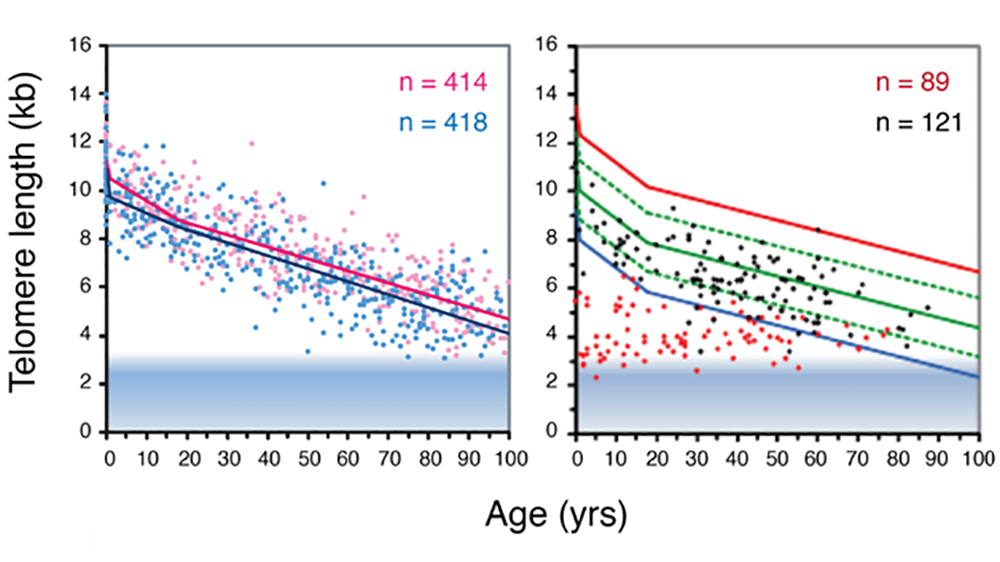
Figure. Telomere length decline with age in human lymphocytes in peripheral blood measured using flow FISH. The left panel shows that a difference in average telomere length between females (pink) and males (blue) persist throughout life. The right panel shows the telomere length in heterozygous carriers of a mutation in either the TERC or the TERT gene (red) and their unaffected siblings (black). Blue line: bottom 1 percent of normal telomere length at indicated age. Figure adapted from (2).
References:
- Baerlocher, G.M., Vulto, I., de Jong, G. and Lansdorp, P.M. (2006) Flow cytometry and FISH to measure the average length of telomeres (flow FISH). Nat Protoc, 1, 2365-2376.
- Aubert, G., Baerlocher, G.M., Vulto, I., Poon, S.S. and Lansdorp, P.M. (2012) Collapse of telomere homeostasis in hematopoietic cells caused by heterozygous mutations in telomerase genes. PLoS Genet, 8, e1002696.
- Lansdorp, P.M. (2022) Sex differences in telomere length, lifespan, and embryonic dyskerin levels. Aging Cell, 21, e13614.
- Savage, S.A. (2025) Telomeres and Human Disease. Cold Spring Harb Perspect Biol, 17.
- Lansdorp, P.M. (2022) Telomeres, aging, and cancer: the big picture. Blood, 139, 813-821.
- Lansdorp, P.M. (2022) Telomeres, Telomerase and Cancer. Arch Med Res, 53, 741-746.
13. Proposed the “silent sister” hypothesis.
In our hunt for genes that regulate the length of telomere repeats we had discovered two similar helicase genes: dog-1 in C.elegans (1) and Rtel in mice (2). We proposed similar roles for the helicases encoded by these genes in the replication of guanine-rich DNA: the unwinding of guanine quadruplex (G4) structures. Long poly-guanine tracts in the C.elegans genome are much more abundant than predicted by chance (3), raising further questions about their role in biology. One possibility is that G4 structures, arising from single stranded G-rich DNA during transcription or replication of G-rich DNA, trigger epigenetic differences between sister chromatids as predicted by the “silent sister hypothesis” (4,5).
References:
- Cheung, I., Schertzer, M., Rose, A. and Lansdorp, P.M. (2002) Disruption of dog-1 in Caenorhabditis elegans triggers deletions upstream of guanine-rich DNA. Nat Genet, 31, 405-409.
- Ding, H., Schertzer, M., Wu, X., Gertsenstein, M., Selig, S., Kammori, M., Pourvali, R., Poon, S., Vulto, I., Chavez, E. et al. (2004) Regulation of murine telomere length by Rtel: an essential gene encoding a helicase-like protein. Cell, 117, 873-886.
- Zhao, Y., O'Neil, N.J. and Rose, A.M. (2007) Poly-G/poly-C tracts in the genomes of Caenorhabditis. BMC Genomics, 8, 403.
- Lansdorp, P.M. (2007) Immortal strands? Give me a break. Cell, 129, 1244-1247.
- Lansdorp, P.M., Falconer, E., Tao, J., Brind'Amour, J. and Naumann, U. (2012) Epigenetic differences between sister chromatids? Ann N Y Acad Sci, 1266, 1-6.
14. Discovered that sister chromatids in daughter cells can be distinguished by inherited parental DNA template strands.
In order to test the silent sister hypothesis (1), we needed to distinguish sister chromatids in paired daughter cells. The solution we initially found was based on the Chromosome Orientation (CO-FISH) technique developed by Ed Goodwin and colleagues in the 90’s (2). Specifically, we found that sister chromatids of mouse chromosomes could be distinguished using PNA probes specific for the large unidirectional arrays of major satellite sequences in murine chromosomes (3).
References:
- Lansdorp, P.M., Falconer, E., Tao, J., Brind'Amour, J. and Naumann, U. (2012) Epigenetic differences between sister chromatids? Ann N Y Acad Sci, 1266, 1-6.
- Goodwin, E. and Meyne, J. (1993) Strand-specific FISH reveals orientation of chromosome 18 alphoid DNA. Cytogenet Cell Genet, 63, 126-127.
- Falconer, E., Chavez, E.A., Henderson, A., Poon, S.S., McKinney, S., Brown, L., Huntsman, D.G. and Lansdorp, P.M. (2010) Identification of sister chromatids by DNA template strand sequences. Nature, 463, 93-97.
15. Developed the single cell Strand-seq method.
Encouraged by the results described in our Nature paper (1), we embarked on developing the single cell DNA template strand sequencing (Strand-seq) technique (2). At the time, we could not have predicted the wide range of applications that this technique would have outside our initial goal: to distinguish sister chromatids in order to test the “silent sister” hypothesis. Strand-seq results are illustrated in the Figure and in a recent review (3).
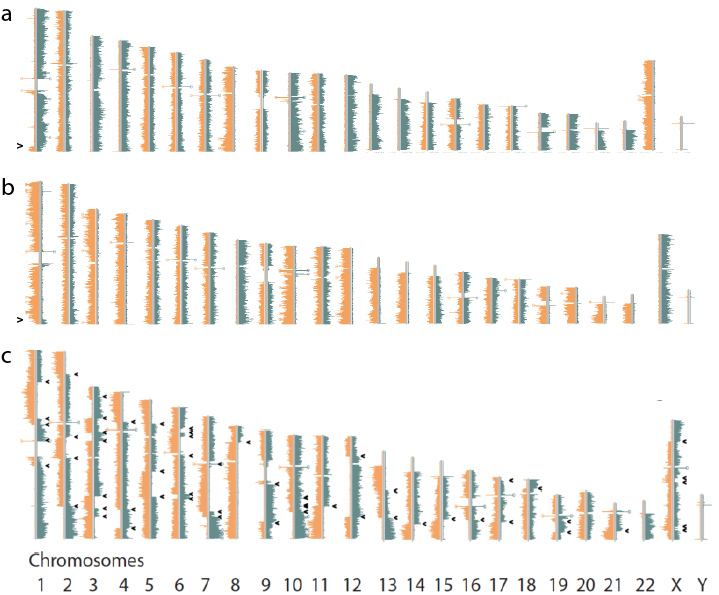
Figure. Strand-seq allows identification and analysis of sister chromatids. The two daughter cells (a and b) of a single parental CD34+ cell show perfect, complementary segregation of parental DNA template strands and only a single sister chromatid exchange event (arrow, bottom chr 1). Note that all chromosomes showing reads mapping to both strands of the reference genome (Chr 2, 5, 6, 7, 9, 11, 14, 16, 17 and 18 in this cell pair) can be used to map SNPs to parental genomes along entire chromosomes and assemble complete physical maps of parental haplotypes. c) Sister chromatid exchange events are very common in cells from patients with Bloom’s syndrome. In this cell 56 SCE events are detected (arrows).
References:
- Falconer, E., Chavez, E.A., Henderson, A., Poon, S.S., McKinney, S., Brown, L., Huntsman, D.G. and Lansdorp, P.M. (2010) Identification of sister chromatids by DNA template strand sequences. Nature, 463, 93-97.
- Falconer, E., Hills, M., Naumann, U., Poon, S.S., Chavez, E.A., Sanders, A.D., Zhao, Y., Hirst, M. and Lansdorp, P.M. (2012) DNA template strand sequencing of single-cells maps genomic rearrangements at high resolution. Nat Methods, 9, 1107-1112.
- Hanlon, V.C.T. and Lansdorp, P.M. (2026) Strand-seq and the future of personalized genomics. Nat Genet.
16. Generated comprehensive maps of human polymorphic inversions.
My lab generated comprehensive maps of polymorphic inversions in the human genome. This work was spearheaded by three graduate students: Ashley Sanders, David Porubsky and Vincent Hanlon. The Figure shows examples of results. Following Ashley’s initial papers (1,2), David reported on inversions and genome instability (3) and Vincent developed software to visualize phased inversions in a genome browser (4).
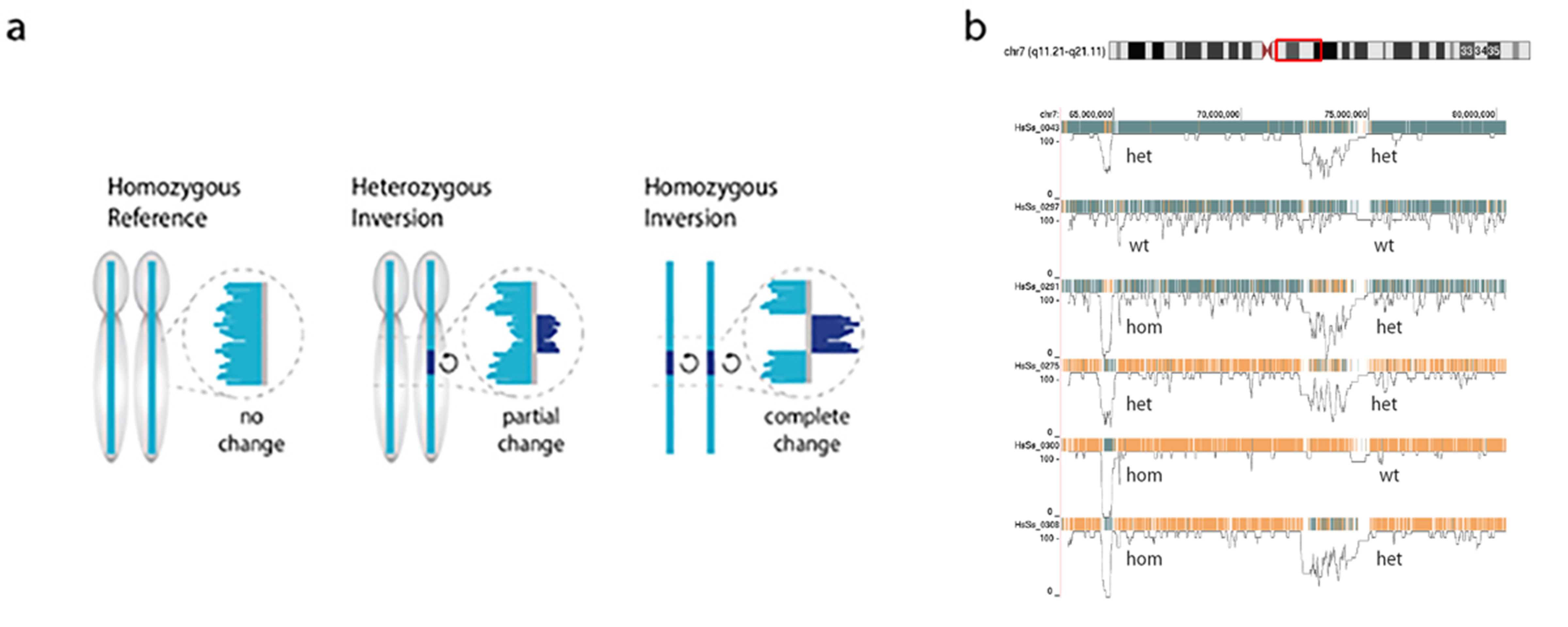
Figure. Sequence reads generated by Strand-seq map either concordant with the human reference genome sequence (a, left) or map to both strands of the reference genome (reflecting a heterozygous inversion (a, middle panel) or completely to the opposite strand of the reference genome reflecting a homozygous inversion (a, right panel). Polymorphic inversions in the human genome generate a large amount of genetic diversity between individuals as is shown in b for a 20Mb segment of chr 7 (red box) for 6 individuals. Note that the role of this type of genetic diversity is largely unexplored because of the difficulty to generate genome-wide information about polymorphic inversions using other sequence modalities.
References:
- Sanders, A.D., Hills, M., Porubsky, D., Guryev, V., Falconer, E. and Lansdorp, P.M. (2016) Characterizing polymorphic inversions in human genomes by single-cell sequencing. Genome Res, 26, 1575-1587.
- Chaisson, M.J.P., Sanders, A.D., Zhao, X., Malhotra, A., Porubsky, D., Rausch, T., Gardner, E.J., Rodriguez, O.L., Guo, L., Collins, R.L. et al. (2019) Multi-platform discovery of haplotype-resolved structural variation in human genomes. Nat Commun, 10, 1784.
- Porubsky, D., Hops, W., Ashraf, H., Hsieh, P., Rodriguez-Martin, B., Yilmaz, F., Ebler, J., Hallast, P., Maria Maggiolini, F.A., Harvey, W.T. et al. (2022) Recurrent inversion polymorphisms in humans associate with genetic instability and genomic disorders. Cell, 185, 1986-2005 e1926.
- Hanlon, V.C.T., Mattsson, C.A., Spierings, D.C.J., Guryev, V. and Lansdorp, P.M. (2021) InvertypeR: Bayesian inversion genotyping with Strand-seq data. BMC Genomics, 22, 582.
17. Developed novel ways to establish haplotypes along entire chromosomes.
The diploid nature of the genome is often incompletely represented in “whole genome sequence” analysis, where a genome is typically represented as a set of unphased variants with respect to a reference genome. However, important biological phenomena such as compound heterozygosity and epistatic effects between enhancers and target genes can only be studied when haplotype-resolved genomes are available. Hence a method that can produce dense and accurate chromosome-length haplotypes is highly desirable. Using a well-studied trio from the HapMap consortium we showed that Strand-seq allows for accurate phasing along entire chromosomes (1). The principle of this method and phasing trio family members are shown in Figure 1.
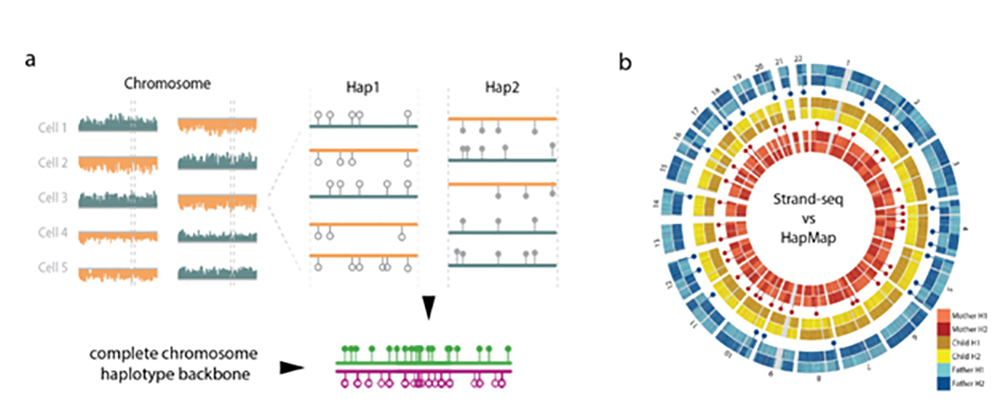
Figure 1. a) Strand-seq provides a physical haplotype map of SNP’s along entire chromosomes. Multiple cells with Strand-seq sequence reads mapping to both strands of the reference genome for a given chromosome (orange and blue) are used to assemble haplotype backbones (1). Such chromosome-long haplotypes correspond accurately (>99%) with HapMap reference data for a child (b, yellow) but not her parents (father blue , mother red), where Strand-seq reveals the location of parental meiotic recombination events (resp. red and blue dots). The haplotype backbones generated by Strand-seq allow complete phasing of human genomes when combined with other type of sequence data (e.g. 10X or PacBio data (1, 2, 3).
When combined with other type of sequencing data, Strand-seq allows near complete phasing (>99% of SNP’s) of human genomes (Fig 2, from 2).
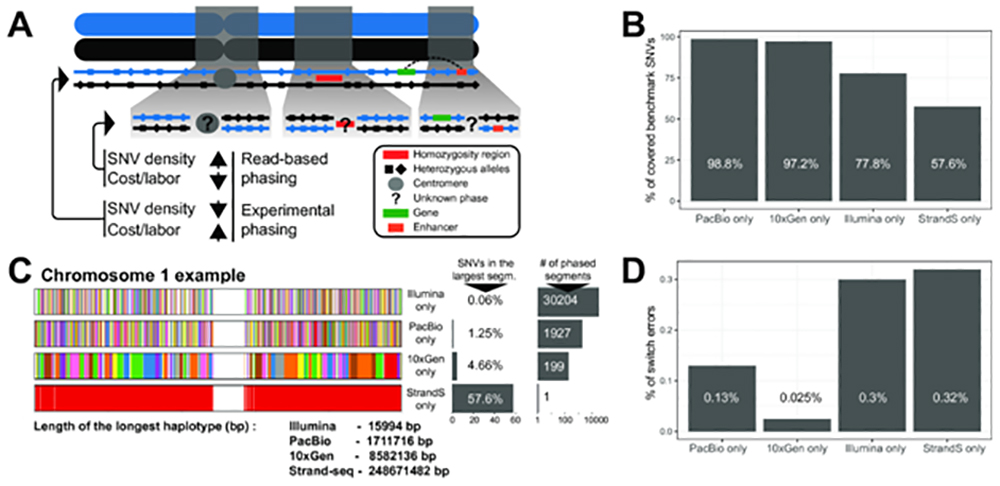
Figure 2. A) Two homologous chromosomes are shown (blue and black). Experimental phasing approaches like Strand-seq can connect heterozygous alleles along whole chromosomes, however, at higher costs (time and labor) and lower density of captured alleles. In contrast, read-based phasing can deliver high-density haplotypes, but only short haplotype segments are assembled with an unknown phase between them. B) Barplot showing the percentage of phased variants, for each sequencing technology, from the total number of reference SNVs (Illumina platinum haplotypes). C) Graphical summary of phased haplotype segments for Illumina, PacBio, 10X Genomics and Strand-seq phasing shown for chromosome 1. Each haplotype segment is colored in a different color with the longest haplotype colored in red. Side bargraph reports the percentage of SNVs phased in the longest haplotype segment. D) Accuracy of each independent phasing approach measured as percentage short switch errors in comparison to benchmark haplotypes.
References:
- Porubsky, D., Sanders, A.D., van Wietmarschen, N., Falconer, E., Hills, M., Spierings, D.C., Bevova, M.R., Guryev, V. and Lansdorp, P.M. (2016) Direct chromosome-length haplotyping by single-cell sequencing. Genome Res, 26, 1565-1574.
- Porubsky, D., Garg, S., Sanders, A.D., Korbel, J.O., Guryev, V., Lansdorp, P.M. and Marschall, T. (2017) Dense and accurate whole-chromosome haplotyping of individual genomes. Nat Commun, 8, 1293.
- Hanlon, V.C.T. and Lansdorp, P.M. (2026) Strand-seq and the future of personalized genomics. Nat Genet.
18. Improved Strand-seq library preparation protocols.
Although Strand-seq data are increasingly in demand, the preparation of single cell Strand-seq libraries is technically challenging (1). Variable results and limited informative reads have hampered widespread application of the Strand-seq method despite its theoretical appeal. We recently reported a simplified “one pot” (OP) method to make Strand-seq libraries that overcomes many of the previous technical challenges (2). Specifically, we developed a nanoliter-volume protocol in which reagents are added cumulatively, DNA purification steps are avoided, and enzymes are inactivated with a thermolabile protease. OP-Strand-seq libraries routinely capture 10%–25% of the genome from a single cell with reduced costs and increased throughput, opening up a wide variety of studies that are being pursued in collaboration with investigators in and outside Vancouver and Canada. A typical result of our current Strand-seq results is shown in the Figure and various issues related to Strand-seq library production are reviewed in (3).
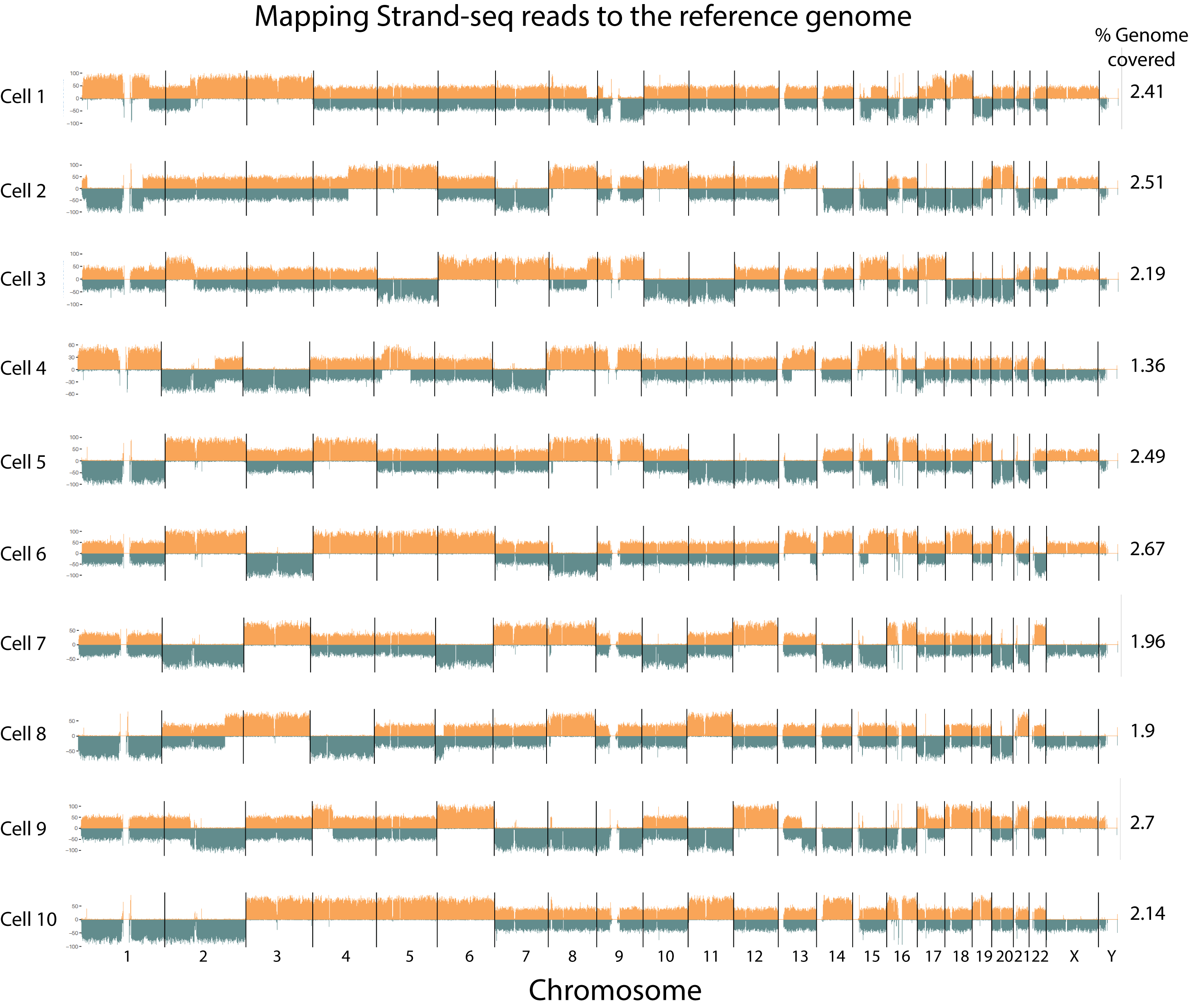
Figure. Example of Strand-seq results (data from 4). Reads mapping to either the plus (+) strand of the human reference genome HG38 (“Crick” Orange) or the minus (-) strand of HG38 (“Watson”, blue) are shown for all chromosomes in single cells from this male (X/Y) individual. Available libraries were not sequenced deeply and the percentage of a haploid genome captured is shown on the right.
References:
- Sanders, A.D., Falconer, E., Hills, M., Spierings, D.C.J. and Lansdorp, P.M. (2017) Single-cell template strand sequencing by Strand-seq enables the characterization of individual homologs. Nat Protoc, 12, 1151-1176.
- Hanlon, V.C.T., Chan, D.D., Hamadeh, Z., Wang, Y., Mattsson, C.A., Spierings, D.C.J., Coope, R.J.N. and Lansdorp, P.M. (2022) Construction of Strand-seq libraries in open nanoliter arrays. Cell Rep Methods, 2, 100150.
- Hanlon, V.C.T. and Lansdorp, P.M. (2026) Strand-seq and the future of personalized genomics. Nat Genet.
- Porubsky, D., Dashnow, H., Sasani, T.A., Logsdon, G.A., Hallast, P., Noyes, M.D., Kronenberg, Z.N., Mokveld, T., Koundinya, N., Nolan, C. et al. (2025) Human de novo mutation rates from a four-generation pedigree reference. Nature, 643, 427-436.
19. Assigning alleles to parent-of-origin chromosomes.
Genetic testing can inform a patient’s inherited risk for developing a disease. However, predicting which side of the family an autosomal variant comes from is often a limitation of current technology. Inability to determine a variant’s parent-of-origin (PofO) (i.e., if it is maternally or paternally inherited) can lead to uncertain patient management and ineffective genetic testing of family members. In collaboration with Intan Schrader, Steven Jones and their collaborators we have developed a protocol using Oxford Nanopore Technologies’ long read sequencing (ONT-LRS) in combination with chromosome-length haplotyping enabled by single cell Strand-seq to allow, for the first time, PofO haplotyping without any sequence information of the parents (1). The principle of this method is shown in the Figure.
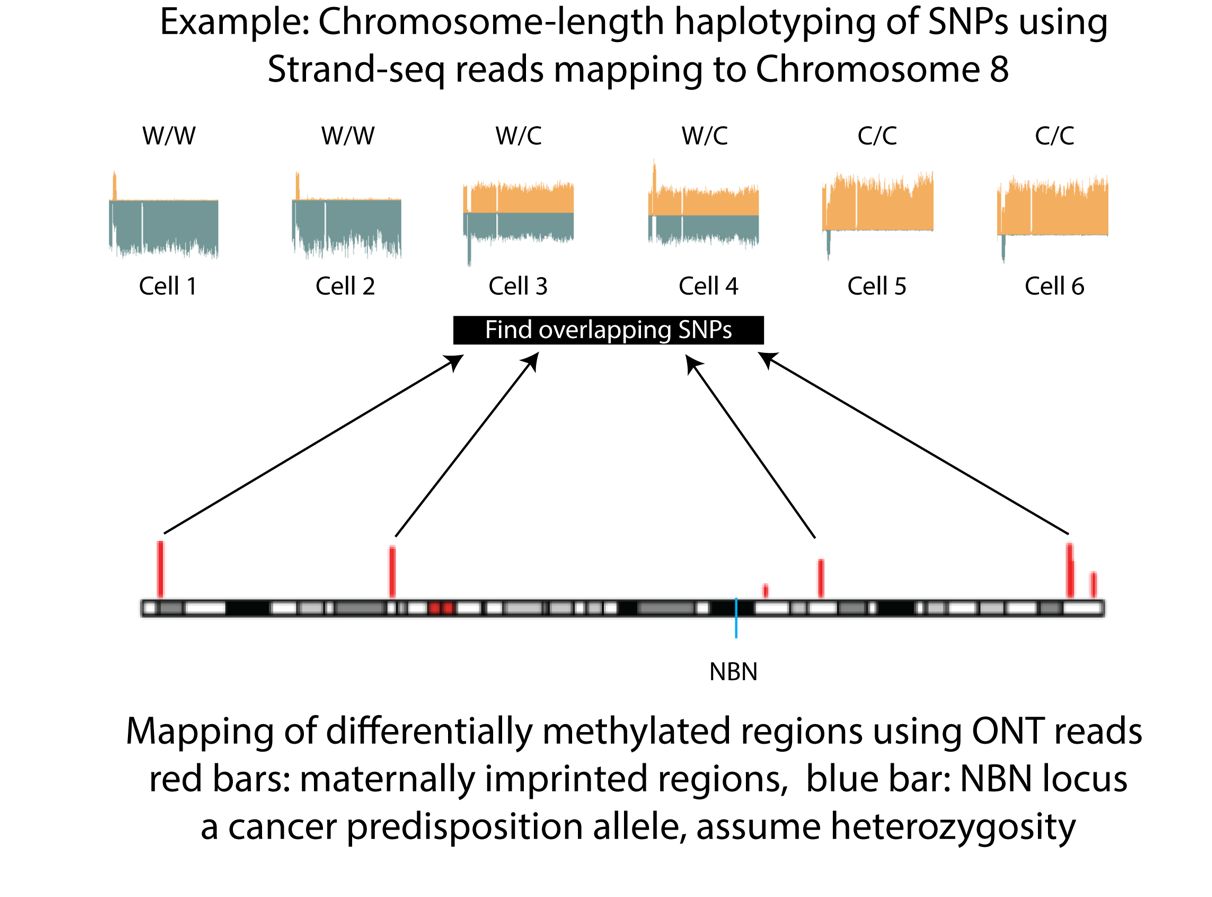
Cells with reads mapping to opposite strands of the reference genome for a particular chromosome occur at a frequency of around 50%. Such cells (3 and 4 in figure) can be used to assign the parent-of-origin haplotype for that chromosome using parental DNA methylation imprints obtained using long read sequencing (1).
References:
- Akbari, V., Hanlon, V.C.T., O’Neill, K., Lefebvre, L., Schrader, K.A., Lansdorp, P.M. and Jones, S.J.M. (2022) Parent-of-origin detection and chromosome-scale haplotyping using long-read DNA methylation sequencing and Strand-seq. Cell Genomics, 100233.
Return to Dr. Lansdorp's profile:
BC Cancer Foundation is the fundraising partner of BC Cancer, which includes BC Cancer Research. Together with our donors, we are changing cancer outcomes for British Columbians by funding innovative research and personalized treatment and care.
